BioTechniques News
Aisha Al-Janabi

New mechanistic insights into the protein complex that powers the bacterial flagellum may assist antibiotic development.
A study led by researchers at the University of Copenhagen (Denmark) used cryo-electron microscopy to investigate how bacteria power their movement. Understanding this cellular mechanism could lead to therapeutic targets for new antibiotics.
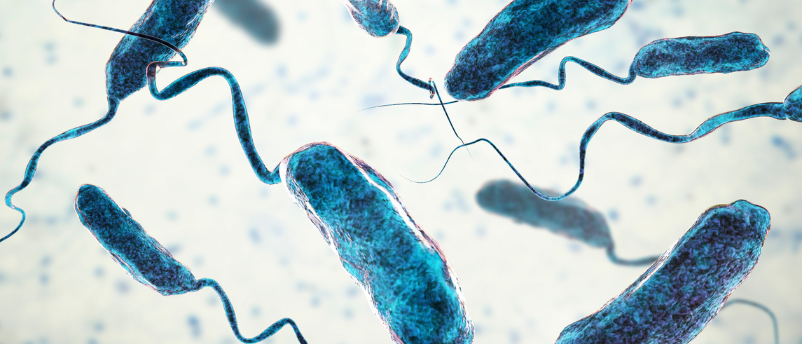
Bacteria move through their environment using flagella, thin filamentous appendages that rotate in a corkscrew-like motion. The rotation of the flagella is powered by stator units called PomAB. When active, the stator units allow specific ions to move across the membrane along an ion gradient and use the energy provided by this process to power the rotation of the rotor and, by extension, the flagellum.
Different species of bacteria contain different forms of motor complexes that use either sodium, potassium or hydrogen ions to power their flagellar motors. However, questions remain regarding the mechanism by which sodium-driven stator units selectively transport sodium ions and not protons or potassium ions, and the mechanism that confers energy from ion transit to motor rotation. These questions are addressed by this new study, which focuses on the bacteria Vibrio alginolyticus, a bacteria that resembles the cholera bacterium.
To obtain cryo-electron microscopy images of PomAB, the researchers first freeze-purified PomAB proteins in thin layers of vitreous ice and imaged them using an electron microscope. By capturing images of PomAB in a variety of orientations, the researchers were able to reconstruct a 3D model representing the structure of PomAB to make inferences about the mechanism of PomAB function.
“The bacteria put a lot of ions on one side of the membrane, and when the ions naturally flow through the small PomAB motor, they allow PomA to rotate around PomB,” explained Nicholas Taylor, the senior author of the study. “Because PomB is anchored to the cell wall and PomA can grab on to the big motor, the flagellar motor, this can rotate the flagellum.”
 Cryo-EM reveals new structural elements in a key ciliary protein complex
Cryo-EM reveals new structural elements in a key ciliary protein complex
Scientists capture a protein complex that transports proteins to cilia in unprecedented detail.
They demonstrated that PomAB adopts an asymmetric pentagon shape, which they predict ensures each stator unit aligns correctly when engaging the rotor. They also characterized regions of PomAB that act as plugs, blocking the path of ions before the rotor is engaged.
The resolution of the data was sufficiently high to discern sodium ions within the structure and the authors utilized computer simulations to plot the path of ions traveling through a cavity in PomAB. They identified amino acid residues that differ from cavities in motors that are driven by other ions. They predicted that these residues play a role in sodium ion specificity and transport.
To confirm their findings, the researchers produced a series of bacterial strains expressing PomAB mutants, in which the identified residues were individually changed, and examined the effect this had on flagella function. They verified that changing any of the identified residues prevented motor function, leaving mutants unable to swim.
Finally, they determined that rotational force transfer is mediated by electrostatic interactions between charged regions on the surface of PomAB and the surface of the rotor.
This study provides new understanding of the mechanisms of PomAB, a process that is crucial for the motility of many bacterial species, including species that cause disease in humans. This makes these motor complexes attractive targets for novel antibiotic design, a vital field of research in the context of emerging antibiotic resistance.
The post Bacterial swimming machinery – a potential antibiotic target? appeared first on BioTechniques.
Powered by WPeMatico
